
For years, the nucleus within a cell was thought to be elastic like a rubber ball, deforming and snapping back into shape as the cell navigated through pores and between fibers inside the human body.
Researchers at Texas A&M University and the University of Florida have discovered that the nucleus is more complex than originally believed, behaving more like a liquid drop than a rubber ball.
“The discovery that the nucleus deforms like a liquid drop calls for a fresh look at how the nuclear shape becomes abnormal in diseases like cancer,” said Dr. Tanmay Lele, Unocal Professor in the Department of Biomedical Engineering.
Lele, a Cancer Prevention & Research Institute of Texas (CPRIT) Scholar, is co-leading the team that uncovered the surprising mechanical behaviors of the nucleus. Their findings were published in Advanced Science in June 2022.
The genetic material governing a cell’s function and behavior, called the genome, is safely stored in the nucleus. Nearly 150 years of looking through microscopes has taught pathologists and researchers that misshapen nuclei are warning signs of diseases like cancer. Cancer cells with such abnormal nuclei are able to migrate to other parts of the body in a process called cancer metastasis, a spreading that can be lethal.
Nuclear shape observations are used in cancer diagnosis even today. But why nuclei become abnormal has remained unclear. Understanding how nuclei become misshapen may help uncover a way to aid cell nuclei in regaining their normal shapes, leading to new approaches for treating cancer.
The findings from this study are critical to understanding how a protective layer surrounding the nucleus, called the lamina, helps preserve nuclear shape while cells crawl through the tortuous paths through pores and around tissue fibers.
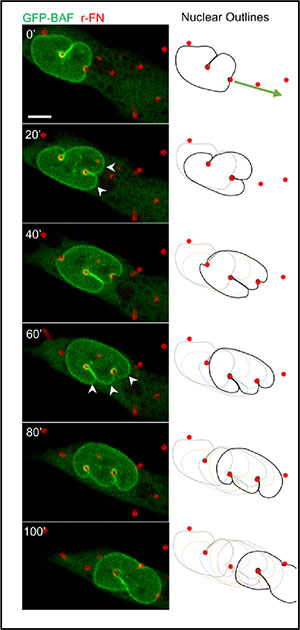
Lele and his fellow researchers began their exploration of nuclear behaviors by placing fibroblasts, the most common type of connective-tissue cells in animals, into a miniature obstacle course of tiny, flexible pillars 1/100th of the width of a human hair. In order for the cells to crawl through this obstacle course, their nuclei had to squeeze in between the pillars. The researchers observed the movements with an advanced high-resolution microscope that could image the 3D shapes of the nuclei.
Imaging revealed that the pillars created deep indentations into the nuclear surface. Yet the overall nuclear shape was preserved, allowing the nucleus to successfully pass like a liquid drop, and unlike a springy elastic rubber ball, through the obstacles.
The research also revealed that a depletion of lamin A/C, one of the normal protein components of lamina, caused the nuclei to get entangled in the obstacles. The discovery suggests that lamin A/C helps maintain the surface tension of the “nuclear drop.”
“Our work points to a fundamental mechanism by which the nucleus preserves its shape and protects its genome,” Lele said. “Our discovery also helps us better understand how misshapen nuclei arise in cancer and how to potentially make them normal again. We are now studying the implications of the drop model for the abnormal nuclear shapes commonly observed in cancer.”
The work is financially supported by a grant from the National Cancer Institute’s Physical Sciences — Oncology Network to Lele, along with additional support from the National Science Foundation to co-principal investigator Dr. Richard B. Dickinson, professor in the Department of Chemical Engineering at the University of Florida. The research is also partially funded by a CPRIT established investigator award to Lele, facilitated through the Texas A&M Engineering Experiment Station.
Aside from Lele and Dickinson, principal investigators and researchers on this project include Drs. Pushkar P. Lele, Cynthia A. Reinhart-King, Kyle J. Roux and Nathan J. Sniadecki. Students include Aditya Katiyar (lead author on the paper), Jian Zhang, Jyot D. Antani, Yifan Yu and Kelsey L. Scott.